Nomenclature and Function
- Standard Name and Systematic Name:
The protein names are based upon the corresponding gene names found on the Locus Summary page (gene naming conventions are outlined in the Gene Nomenclature Guide). Standard protein names are non-italicized, and are derived from gene names by making the first character uppercase, the remaining alphabetical characters lowercase, and appending the lowercase letter "p". For example, the protein encoded by the C. albicans gene ABC1 (systematic name orf19.3331, allele name orf19.10842) has standard name Abc1p, systematic name Orf19.3331p, and allele name Orf19.10842p. See the CGD Sequence Documentation Page for more details. - Description:
A concise summary of the biological role and molecular function of the protein.
Structural Information
Provides:
- A link to a page listing the structurally related proteins identified by a BLASTP search of the RCSB Protein Databank (PDB); please see the PDB Homolog documentation page for more details about this page.
- A link directly to the RCSB Protein Databank (PDB).
- Summary information for the top PDB hit, including a thumbnail diagram and link to the protein alignment.
Conserved Domains
Provides:
- A link to the Domains/Motifs page, which lists conserved features in the protein.
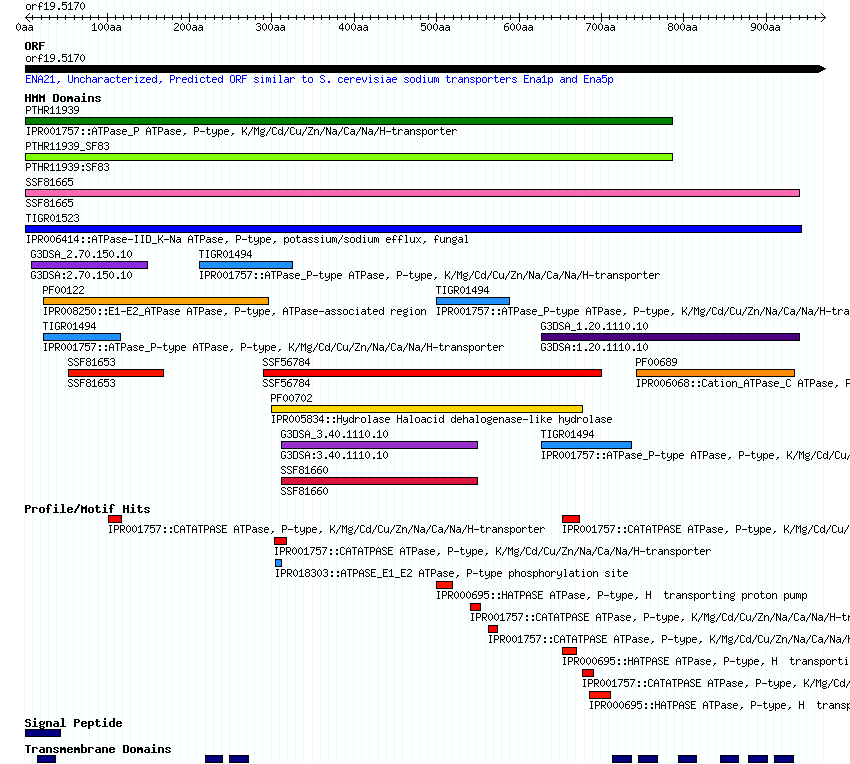
- A thumbnail image and link to the CGD Proteome Browser, which interactively displays conserved domains identified using the protein sequence to query the InterPro database:
Sequence Detail
Summary of the predicted ORF translation, including:
- Length (a.a.): the predicted full length of the translated gene product.
- Molecular Weight (Da): calculated for the predicted full-length sequence.
- A link to the Physicochemical Properties page, which displays other properties and statistics calculated for the sequence.
- Predicted Sequence: the amino acid sequence itself, with a button to download the sequence in FASTA format.
Homologs
Provides links to CGD resources that can be used to identify homologs of the query protein:
- BLAST against other Candida sequences: Runs BLAST locally using protein sequence as query to identify Candida homologs.
External Sequence Databases
Provides links to various other databases containing the protein sequence.
References
This section lists the references used to curate information displayed in the Standard Name, Systematic Name, and Description fields on the page. Note that this section is not a comprehensive listing of publications relevant to this gene. To retrieve a list of all publications annotated to this gene, select the "Literature" tab at the top of the page.
 Return to CGD
Return to CGD